Demo Script
Image creation
A binary image with an ellipsis
>>> f = iagaussian([100,100], [45,50], [[20,5],[5,10]]) > 0.0000001
>>> iashow(f)
(100, 100) Min= 0 Max= 1 Mean=0.098 Std=0.30
 |
|
| f |
Feature vector
The coordinates of each 1 pixel are the features: cols and rows coordinates. The mean value is the centroid.
>>> import Numeric, MLab
>>> (rows, cols) = iaind2sub(f.shape, Numeric.nonzero(Numeric.ravel(f)))
>>> x = Numeric.concatenate((cols[:,Numeric.NewAxis], rows[:,Numeric.NewAxis]), 1)
>>> mx = MLab.mean(x)
>>> print mx
[ 50. 45.]
Computing eigenvalues and eigenvectors
The eigenvalues and eigenvectors are computed from the covariance. The eigenvalues are sorted in decrescent order
>>> Cx = MLab.cov(x)
>>> print Cx
[[ 59.31288344 29.65644172] [ 29.65644172 117.32924335]]
>>> [aval, avec] = MLab.eig(Cx)
>>> aux = Numeric.argsort(aval)[::-1]
>>> aval = MLab.diag(Numeric.take(aval, aux))
>>> print aval
[[ 129.8057478 0. ] [ 0. 46.83637899]]
>>> avec = Numeric.take(avec, aux)
>>> print avec
[[-0.38778193 -0.92175115] [-0.92175115 0.38778193]]
Measure the angle of inclination
The direction of the eigenvector of the largest eigenvalue gives the inclination of the elongated figure
>>> a1 = Numeric.sqrt(aval[0,0])
>>> a2 = Numeric.sqrt(aval[1,1])
>>> vec1 = avec[:,0]
>>> vec2 = avec[:,1]
>>> theta = Numeric.arctan(1.*vec1[1]/vec1[0])*180/Numeric.pi
>>> print 'angle is %3f degrees' % (theta)
angle is 67.183445 degrees
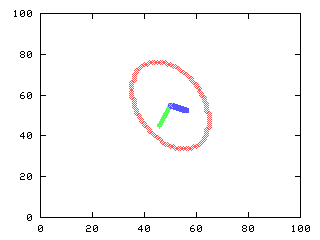
Plot the eigenvectors and centroid
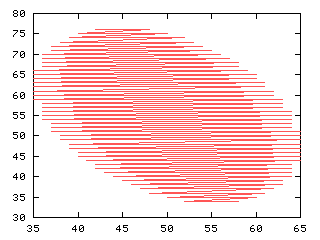
The eigenvectors are placed at the centroid and scaled by the square root of its correspondent eigenvalue
>>> iashow(f)
(100, 100) Min= 0 Max= 1 Mean=0.098 Std=0.30
>>> x_,y_ = iaind2sub(f.shape, Numeric.nonzero(Numeric.ravel(iacontour(f))))
>>> g,d0 = iaplot(y_, 100-x_)
>>> # esboça os autovetores na imagem
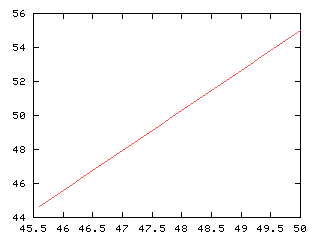
>>> x1, y1 = Numeric.arange(mx[0], 100-mx[0]+a1*vec1[0], -0.1), Numeric.arange(mx[1], mx[1]-a1*vec1[1],0.235)
>>> g,d1 = iaplot(x1, 100-y1) # largest in green
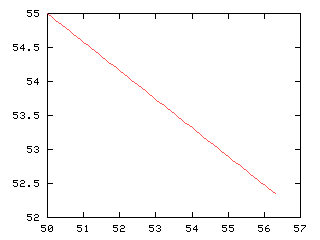
>>> x2, y2 = Numeric.arange(mx[0], mx[0]-a2*vec2[0], 0.1), Numeric.arange(mx[1], mx[1]+a2*vec2[1],0.042)
>>> g,d2 = iaplot(x2, 100-y2) # smaller in blue
>>> g,d3 = iaplot(mx[0], 100-mx[1]) # centroid in magenta
>>> g('set data style points')
>>> g('set xrange [0:100]')
>>> g('set yrange [0:100]')
>>> g.plot(d0, d1, d2, d3)
>>>
 |
 |
|
| f | y_, 100-x_ |
 |
 |
|
| x1, 100-y1 | x2, 100-y2 |
 |
 |
|
| mx[0], 100-mx[1] | d0, d1, d2, d3 |
Compute the Hotelling transform
The Hotelling transform, also called Karhunen-Loeve (K-L) transform, or the method of principal components, is computed below
>>> y = Numeric.transpose(Numeric.matrixmultiply(avec, Numeric.transpose(x-mx)))
>>> my = MLab.mean(y)
>>> print my
[ 8.34651835e-017 -2.23178643e-016]
>>> Cy = MLab.cov(y)
>>> print Cy
[[ 1.29805748e+002 -5.53976315e-015] [-5.53976315e-015 4.68363790e+001]]
>>> print Numeric.floor(0.5 + Numeric.sqrt(Cy))
math domain error
Display the transformed data
The centroid of the transformed data is zero (0,0). To visualize it as an image, the features are translated by the centroid of the original data, so that only the rotation effect of the Hotelling transform is visualized.
>>> ytrans = Numeric.floor(0.5 + y + Numeric.resize(mx, (x.shape[0], 2)))
>>> g = Numeric.zeros(f.shape)
>>> i = iasub2ind(f.shape, ytrans[:,1], ytrans[:,0])
>>> Numeric.put(g, i, 1)
>>> iashow(g)
(100, 100) Min= 0 Max= 1 Mean=0.094 Std=0.29
 |
|
| g |
Image read and display
The RGB color image is read and displayed
>>> f = iaread('boat.ppm')
>>> iashow(f)
(3, 257, 256) Min= 0 Max= 218 Mean=94.533 Std=57.35
 |
|
| f |
Extracting and display RGB components
The color components are stored in the third dimension of the image array
Feature vector: R, G and B values
The features are the red, green and blue components. The mean vector is the average color in the image. The eigenvalues and eigenvectors are computed. The dimension of the covariance matrix is 3x3 as there are 3 features in use
>>> x = 1.*Numeric.concatenate((Numeric.ravel(r)[:,Numeric.NewAxis], Numeric.ravel(g)[:,Numeric.NewAxis], Numeric.ravel(b)[:,Numeric.NewAxis]), 1)
>>> mx = MLab.mean(x)
>>> print mx
[ 101.26770732 94.9191999 87.41316573]
>>> Cx = MLab.cov(x)
>>> print Cx
[[ 3497.95108557 3273.20413327 3086.73663437] [ 3273.20413327 3239.67139072 3103.11181452] [ 3086.73663437 3103.11181452 3032.39588874]]
>>> [aval, avec] = MLab.eig(Cx)
>>> aux = Numeric.argsort(aval)[::-1]
>>> aval = MLab.diag(Numeric.take(aval, aux))
>>> print aval
[[ 9572.66965395 0. 0. ] [ 0. 180.2290775 0. ] [ 0. 0. 17.11963358]]
>>> avec = Numeric.take(avec, aux)
>>> print avec
[[-0.59515464 -0.58010086 -0.55612404] [-0.76785348 0.20637147 0.60647494] [ 0.2370485 -0.78796815 0.5682554 ]]
Hotelling transform
The K-L transform is computed. The mean vector is zero and the covariance matrix is decorrelated. We can see the values of the standard deviation of the first, second and third components
>>> y = Numeric.transpose(Numeric.matrixmultiply(avec, Numeric.transpose(x-mx)))
>>> my = MLab.mean(y)
>>> print my
[ 1.86724609e-013 9.30065579e-015 1.03426779e-014]
>>> Cy = MLab.cov(y)
>>> print Cy
[[ 9.57266965e+003 -8.78852081e-011 -7.44802346e-011] [-8.78852081e-011 1.80229077e+002 -7.44824028e-012] [-7.44802346e-011 -7.44824028e-012 1.71196336e+001]]
>>> Cy = Cy * (1 - (-1e-10 < Cy < 1e-10))
>>> print Numeric.floor(0.5 + Numeric.sqrt(Cy))
[[ 98. 0. 0.] [ 0. 13. 0.] [ 0. 0. 4.]]
Displaying each component of the transformed image
The transformed features are put back in three different images with the g1 with the first component (larger variance) and g3 with the smaller variance
>>> g1 = y[:,0]
>>> g2 = y[:,1]
>>> g3 = y[:,2]
>>> g1 = Numeric.reshape(g1, r.shape)
>>> g2 = Numeric.reshape(g2, r.shape)
>>> g3 = Numeric.reshape(g3, r.shape)
>>> iashow(g1)
(257, 256) Min= -194.702376045 Max= 163.945218189 Mean=0.000 Std=97.84
>>> iashow(g2)
(257, 256) Min= -53.4192670921 Max= 81.6627408632 Mean=0.000 Std=13.42
>>> iashow(g3)
(257, 256) Min= -17.361967919 Max= 20.538452635 Mean=0.000 Std=4.14
 |
 |
|
| g1 | g2 |
 |
|
| g3 |


